 Genome - The Autobiography of a Species in 23 Chapters by Matt Ridley came highly recommended on Amazon. Despite the fact that the book was published in 2000, I decided to read it. The
structure of the book is rather clever: each chapter focuses on one
chromosome. This format
allows Ridley to expand the metaphor that the human genome is a book:
each chromosome is a chapter; the genes are the sentences; the codons
(the three letter DNA sequences that the cellular machinery reads) are
the words; and the DNA nucleotides are the letters. This kind of metaphor can help non-scientists visualize and remember the
organization of the genome. Unfortunately, Ridley eschews the proper scientific
terminology, calling codons "words" and
nucleotides "letters". Science writing should
teach people more about the subject matter; to eliminate the use of fundamental terminology seems a folly.
Genome - The Autobiography of a Species in 23 Chapters by Matt Ridley came highly recommended on Amazon. Despite the fact that the book was published in 2000, I decided to read it. The
structure of the book is rather clever: each chapter focuses on one
chromosome. This format
allows Ridley to expand the metaphor that the human genome is a book:
each chromosome is a chapter; the genes are the sentences; the codons
(the three letter DNA sequences that the cellular machinery reads) are
the words; and the DNA nucleotides are the letters. This kind of metaphor can help non-scientists visualize and remember the
organization of the genome. Unfortunately, Ridley eschews the proper scientific
terminology, calling codons "words" and
nucleotides "letters". Science writing should
teach people more about the subject matter; to eliminate the use of fundamental terminology seems a folly.
In each chapter, the author highlights one gene of interest on the
chromosome. For example, the chapter on chromosome 13 introduces BRCA2. Like
the Jeff Wheelwright book I reviewed previously, Ridley discusses the population genetics of BRCA2. In some chapters, he writes about genetic lessons
from a particular chromosome. In some cases, the conceit works brilliantly (e.g., the chapter on the X and Y chromosomes examines sexually antagonistic genes), but other times, it was not as successful. Ridley uses
Chapter 21 to discuss eugenics. While the history of the topic is interesting,
the connection to chromosome 21 seems a bit tenuous. Parents are increasingly using
prenatal screening to detect chromosomal abnormalities, the most common of
which is Down syndrome, which is caused by an extra copy of chromosome 21. I
found the book worked best when there was an obvious candidate gene to discuss
on the chromosome.
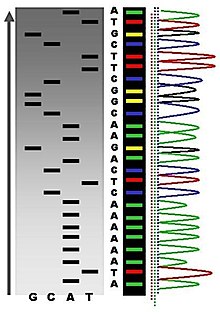 It is amazing to consider how much the field of genomics has changed since the publication of Ridley's book (February 2000). In June of that year, the initial rough draft of the human genome was published. When the sequence was officially completed in April 2003, the final cost was estimated to be $2.7 billion; the project took more than a decade. Today, we are approaching the benchmark of the $1000 genome (discussed here and here). The next step is to improve the speed and portability of sequencing equipment as well as the ability to process and analyze the data.
It is amazing to consider how much the field of genomics has changed since the publication of Ridley's book (February 2000). In June of that year, the initial rough draft of the human genome was published. When the sequence was officially completed in April 2003, the final cost was estimated to be $2.7 billion; the project took more than a decade. Today, we are approaching the benchmark of the $1000 genome (discussed here and here). The next step is to improve the speed and portability of sequencing equipment as well as the ability to process and analyze the data.
The human genome project has been considered a success in terms of yields for basic research. However, there has been some disappointment that the completion of the project hasn't led to more clinical applications (for more specifics, see these editorials from Francis Collins and Craig Venter, the heads of the Human Genome Project). One problem is that very few diseases are caused by a single gene. Another confounding factor is that there is 1-3% difference between any two individuals' genome sequences. These variations can complicate genomic analyses. To perform a genomic study of a population with a genetic
condition, knowing the differences that are normally present in the
genome can help narrow down the possible regions that are linked to the
condition of interest. Thus, having more complete sequences available will allow scientists to connect DNA sequences with genetic conditions. The 1000 Genomes Project plans to identify all the genetic variations present in the human population. The initial phase of this project was completed in just over four years with the publication of 1,092 complete human genome sequences from 14 populations across the globe. In the coming years, the project plans to complete a total of 2,500 genome sequences to improve the representation of various human populations across the globe.
Another major change in the genomic landscape is the advent of personal genomics, such as 23andme. These companies offer DNA analysis service that supplies information on your possible ancestry as well as information about other genetic markers. These markers include both the innocuous ones, like whether you can detect a terrible smell in your urine after eating asparagus, and genes linked to possible health risks (e.g., BRCA, Huntington's disease). After intervention by the FDA, the service is now limited solely to ancestry. Scientific American had a fascinating piece about why we should really be concerned about companies like 23andme (TLDR: they are collecting and storing your most personal data
– your DNA).
--------------------------------------------------------------------------------------------------------
Want read more?
* The Human Genome at 10 Special Issue in Nature covers the changes in the genomics landscape with editorials and articles by a number of major players in the field.
* There is also abundant information on other "big science" approaches to genomics, such as the HapMap, the ENCODE project, and The Cancer Genome Atlas [TCGA]. I have not explored these for the sake of brevity.


